Platform
Creating exceptional medicines with structure and ReSOLVE®
Our proprietary drug discovery platform ReSOLVE® combines the latest advances in AI/ML, protein science, structural biology, and biophysics, to substantially increase the speed and accuracy of small-molecule drug discovery.
Now available: the Rotamer And Protonation Assignment (RAPA) tool
Enables the generation of more accurate protein models that cover the full
set of possible configurations of a protein
Find it on GitHub
Proteins are constantly changing shape, forming new pockets, or transforming the contours of existing pockets. In a cell, proteins are surrounded by water, which fills the pockets of the protein as it changes shape. When interacting with a protein binding pocket, potent small molecules displace this water.
ReSOLVE® is the only platform that can: model all conformations of a protein, identify druggable pockets, understand the dynamic water networks of a pocket, and use those water networks to generate a blueprint for small molecule therapeutics, which we refer to as a hydrocophore®. With ReSOLVE®, we are able to identify novel molecules for targets that have been inaccessible by small molecules and quickly optimize these molecules towards a development candidate.
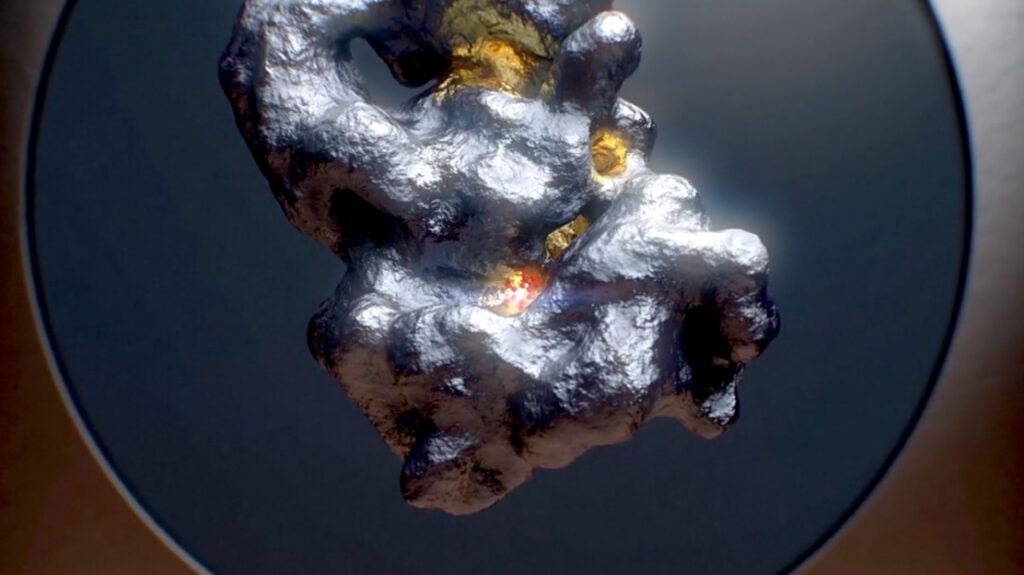

The HYDROCOPHORE®:
a “blueprint” for small molecule design
With ReSOLVE®, we can extract the dynamic water structure of a protein’s binding pockets (the hydrocophore®) and use it as a blueprint for the identification and optimization of small molecule binders. ReSOLVE® allows us to understand the shape, polarity, and potential druggability of binding pockets at a new level of resolution, driving unparalleled accuracy in virtual screening.
The hydrocophore® enables rapid virtual screening of libraries containing billions of compounds, efficiently identifying potent and structurally unique chemical matter.
We restrict the virtual screen to only pursue chemical matter resembling the hydrocophore®, thus generating far fewer virtual hits for wet-lab validation than traditional methods. Reducing the number of compounds that are synthesized and tested greatly reduces the cost and time needed for each screening iteration. Validated screening hits along with the hydrocophore® are used to drive additional screening and further optimization of chemical matter, as demonstrated by our cGAS program. We optimize only promising chemical matter, further improving our overall efficiency. In the time needed for a traditional screening process to develop one development candidate for one target, ReSOLVE® enables the generation of development candidates for multiple different targets, allowing simultaneous preclinical testing of multiple targets and efficient identification of the most promising targets for a particular disease.
Efficient virtual screens that generate NOVEL CHEMICAL MATTER
ReSOLVE® in Action – VENT-03
VENT-03 is the first cGAS inhibitor to enter clinical development. VENT-03 is also Ventus’ first ReSOLVE® success story.
Through the application of ReSOLVE®, we found that the catalytic pocket of cGAS adopts many conformations that had not been previously described. Ventus uniquely developed a detailed understanding of the solvation structure of cGAS and identified previously unknown conformations of the active site. Importantly, we found that some conformations were significantly larger than those observed experimentally, offering new opportunities for optimization. These proprietary insights enabled the development of chemical matter with high potency and desirable pharmacological properties, leading to our portfolio of orally bioavailable small-molecule cGAS inhibitors, including VENT-03.
A new level of resolution with ReSOLVE®
Protein science expertise:
Highly enabled structure-based
drug design
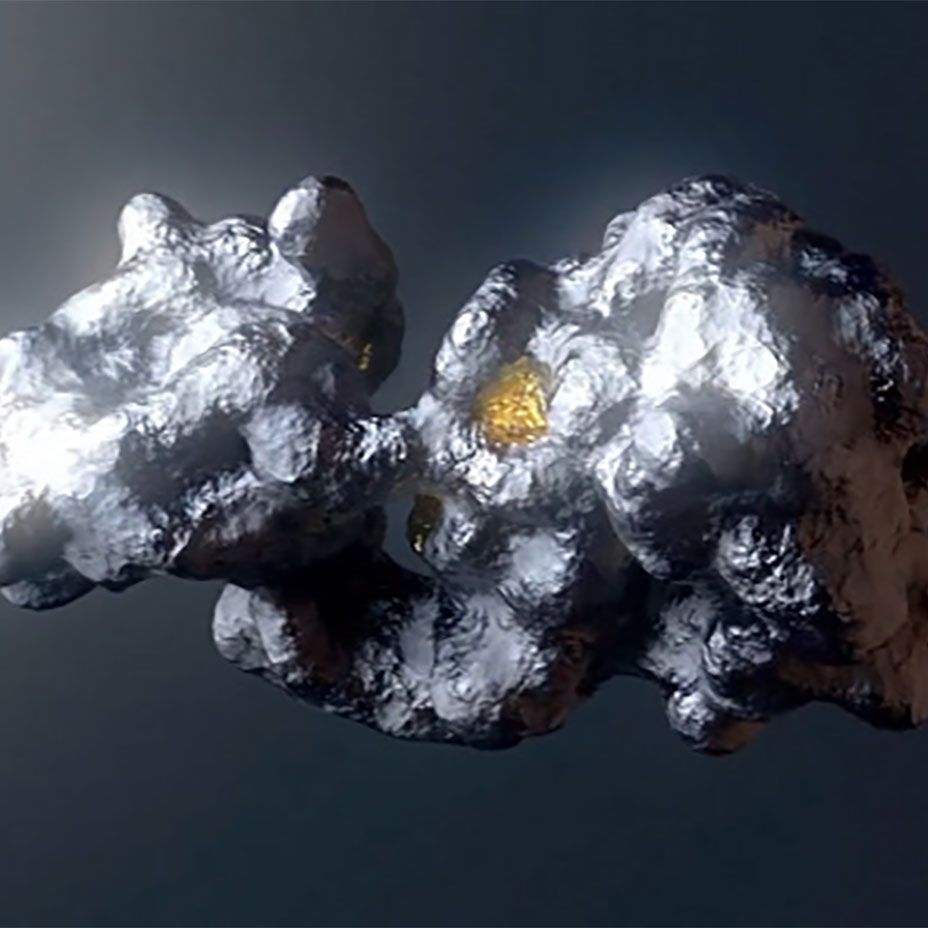

Unrestrained molecular dynamics: Modeling and clustering a moving protein’s many conformations
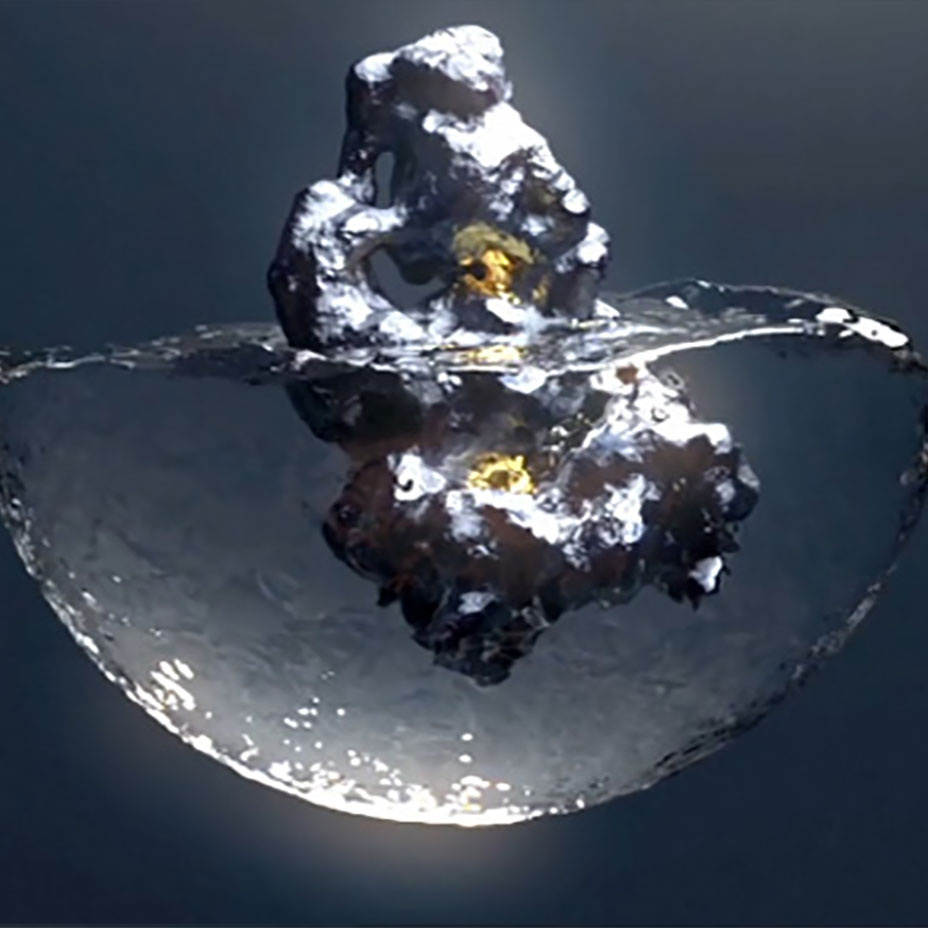

Accurate modeling of the
dynamic solvation structure of
binding pockets
Protein science expertise:
Highly enabled structure-based
drug design
High quality protein target structural information is required to fully take advantage of our unique unrestrained molecular dynamics and solvation analysis tools.
Ventus has deep in-house expertise in protein engineering, expression, and structure determination. This has allowed us to solve high-resolution structures of challenging targets, such as NLRP3 and cGAS, and proceed to structure-based drug design.
Unrestrained molecular dynamics: Modeling and clustering a moving protein’s many conformations
Computer simulations of proteins in motion produce millions of conformations for evaluation. Many existing conformation clustering methods result in configurations that don’t exist in nature and will lead to a dead-end in the drug discovery process.
ReSOLVE® addresses this limitation by enabling the unbiased modeling of a broad set of configurations for a dynamic protein, including those that are rare.
Accurate modeling of the
dynamic solvation structure of
binding pockets
Traditional methods to identify small molecules that can bind to pockets on proteins are not designed to account for the importance of the dynamic nature of the solvation structure within a binding pocket.
ReSOLVE® addresses this limitation by modeling the dynamic solvation structure of each conformation to identify a hydrocophore®, a three-dimensional blueprint of an existing pocket.
Partnering opportunities
ReSOLVE® can screen targets that are outside of scope for Ventus. If you are interested in learning more about ReSOLVE®or collaboration opportunities, please contact us by filling out the form here.

